Document Type : Short Communication
Author
Department of Chemical Sciences, Adekunle Ajasin University, Akungba-Akoko, Nigeria
Abstract
Drug resistance by persisters is a global issue that requires urgent attention. Quinolines and quinolones have proven to be important cores in several bioactive organic chemotypes, based on this, the antimicrobial activity of six derivatives of 2,4-diarylquinolines was evaluated using the disc diffusion method at 5-20 µM/mL. The bioactivity evaluation was conducted on four bacteria examples, two of which are ESKAPE bacteria (Staphylococcus aureus, Streptococcus pneumonia, Escherichia coli and Pseudomonas aeruginosa). The results indicated promising minimum inhibitory concentrations (5 and 10 µM/mL in some cases) and encouraging zones of inhibition against the 4 germs while interesting and reasonable structure-activity relationships were established at the end of the study.
Graphical Abstract
Keywords
Introduction
The attainability of Millennium Development Goals (MDGs) and Sustainable Development Goals (SDGs) is threatened by antimicrobial resistance, antibiotic persistence and tolerance by bacteria [1]. Antimicrobial resistance, antibiotic persistence and tolerance by persisters have been implicated as the cause of treatment failure in patients receiving antibiotics [2].
Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter spp are represented by the acronym ESKAPE. These pathogens have shown resistance to current antibiotics [3]. ESKAPE germs possess resistance traits caused by mechanical protection as a result of their ability to synthesis biofilm among other attributes [4]. Therefore, there is a need for continual search and development of novel antibiotics while taking other measures such as combination of antibiotics with adjuvants, photodynamic and photochemical therapies [3].
Quinoline and quinolone scaffolds are broad spectrum cores in terms of bioactivities while some of the current antibiotics have quinoline/quinolone cores [5-7]. For example, Ciprofloxacin 1 (Figure 1), a quinolone, is a quinolin-4(1H) core and its bioactivity against bacteria relies on the inhibition of DNA gyrase and topoisomerase IV [8]. Clioquinol 2 (Figure 1) on the other hand has a quinoline core, it is a dihalogenated 8-hydroxyquinoline. It has antiseptic activity, a useful drug against drug-resistant Candida and it acts by inhibiting the development of hyphal which in turn block the formation of biofilm [9, 10]. Recently, the activity of 2,4-diarylquinoline derivatives 3 (Figure 1) against Trypanosoma brucei brucei was reported [11].
Figure 1. Structural representations of quinolone and quinoline-based antibiotics and 2,4-diarylquonoline
As a contribution to the development of new and efficient antimicrobial chemotypes which may help overcome the problem of antimicrobial resistance, especially in ESKAPE bacteria, this short communication further explores the application of derivatives of 2,4-diarylquinolines (compounds 3) as antimicrobial agents. The quinoline derivatives used for this study were available from previous study [11] while the unavailable ones were freshly prepared. The molecules were tested against Staphylococcus aureus, Streptococcus pneumonia, Escherichia coli and Pseudomonas aeruginosa. It should be noted that this group of compounds are not cytotoxic having previously screened them for toxicity against HeLa cells [11].
Experimental
Materials, chemicals and bacterial strains
All the chemicals and solvents were purchased from the Sigma Aldrich and used as supplied.
The four bacteria (S. aureus, S. pneumonia, E. coli and P. aeruginosa) used for this study were provided by the Microbiology laboratory of Lagos State Teaching Hospital (LUTH), Nigeria. The isolates were inoculated on the Ageller Mueller-Hinton culture medium.
Antibacterial test
Test for susceptibility of bacteria to antibiotics was carried out using the agar disc diffusion assay [12]. This method essentially involves the spread of the 24-h bacterial culture on the surface of the Mueller-Hinton agar plate. Different concentrations of 20, 10, and 5 µM solutions of the molecules under study were prepared by serial dilution. A disc of sterile 6-mm filter paper was saturated with the different concentration of solution of the studied compounds in dimethyl sulfoxide (DMSO) and allowed to diffuse for 1h. The Petri dishes were incubated at 37 °C for 24 h and the diameter of the zone of inhibition was taken. The measured diameter was compared to that of Ofloxacin 4 standard (Figure 2).
Figure 2. Ofloxacin, a standard antibiotic
Results and Discussion
Chemistry
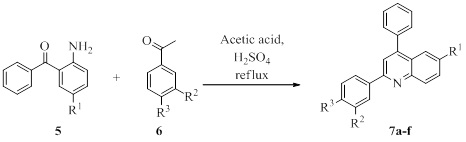
The 2,4-Diarylquinolines (7a-f) were prepared from 2-aminobenzophenones (5) and acetophenones (6) as described by Oluwafemi et al (Scheme 1) [11]. The desired products were obtained in essentially pure form as evidenced by thin layer chromatography (TLC) and nuclear magnetic resonance (NMR) spectroscopy [11].
Scheme 1. Synthesis of 2,4-Diarylquinolines
Antimicrobial activity
The structure-activity relationships observed for 6-chloro-2,4-diphenylquinoline 7a; 6-chloro-2-(4-chlorophenyl)-4-phenylquinoline 7b; 2-(4-bromophenyl)-6-chloro-4-phenylquinoline 7c; 6-chloro-2-[3-(trifluoromethyl)phenyl]-4-phenylquinoline 7d; 6-chloro-2-[4-(trifluoromethyl)phenyl]-4-phenylquinoline 7e and 6-nitro-2-[4-(trifluoromethyl)phenyl]-4-phenylquinoline 7f after screening against S. aureus, S. pneumonia, E. coli and P. aeruginosa is summarised in Table 1. Compound 7a gave minimum inhibitory concentrations (MIC) of 10 µM/ml when tested on S. pneumonia, E. coli and P. aeruginosa while 20 µM/mL was recorded for S. aureus. With chlorine as R1 and R3 in compound 7b, the MICs were 20 and 10 µM/mL, respectively for S. aureus and S. pneumonia while 7b was not active against E. coli and P. aeruguinosa. Changing R3 to bromine to have 7c, there was activity against all the microbs (MICs of 10 µM/ml against S. aureus, S. pneumonia and P. aeruguinosa but a MIC of 20 µM/mL was recorded for E. coli). Compound 7c was very good against S. aureus because 25 mm was obtained as the zone of inhibition.
Table 1. The minimum inhibitory concentration of compounds 7a-f against S. aureus, S. pneumonia, E. coli and P. aeruguinosa
On exploring trifluromethane functionalized 2,4-diarylquinolines (in compounds 7d-f), It was found that changing R2 to trifluormethane (-CF3) to obtain 7d, the compound was active against the four microbs, but on moving the –CF3 functionality to R3 position (compound 7e), the bioactivity disappeared. When the R1 in 7e was changed to a nitro group (-NO2) to obtain 7f, bioactivity was experienced with MICs of 5 µM/mL against S. aureus (zone of inhibition = 15 mm), 10 µM/mL against S. pneumonia but 20 µM/mL against E. coli and P. aeruguinosa (zone of inhibition = 14 and 19 mm, respectively). The improved bioactivity could be attributed to the presence of the strongly electron withdrawing nitro group as R1.
Conclusion
In this research study, six 2,-4-diarylquinolines were screened for bacteria susceptibility against S. aureus, S. pneumonia, E. coli and P. aeruginosa at concentrations of 5-20 µM/mL. Results obtained established the importance of the nitro-, bromo-, chloro- and trifluoromethyl- substituents on the 2,4-diarylquinoline core in relation to bioactivity against microbs. This work also established that 2,4-diarylquinolines, especially the 2-(4-bromophenyl)-6-chloro-4-phenylquinoline 7c and 6-nitro-2-[4-(trifluoromethyl)phenyl]-4-phenylquinoline 7f are suitable to be considered for a prospective hit-to-lead expansion in a future pharmacomodulation process as search for better antibiotics against ESKAPE pathogens continues. It should be noted that the mode of inhibitory action/mechanism of antibiotic action of these compounds against S. aureus, S. pneumonia, E. coli and P. aeruginosa are not known at the moment.
Acknowledgement
The author is indebted to the Departments of Chemical Sciences and Microbiology, Adekunle Ajasin University, Akungba-Akoko, Nigeria.
Disclosure statement
No potential conflict of interest was reported by the author.


.jpg)
.jpg)