Document Type : Original Research Article
Authors
- Khaled Muftah Elsherif 1
- Hasan Aberrwaila 2
- Abdulaziz Suliman Aburowais 3
- Ali Mansour Etwaish 2
- Ali Meftah Elassawi 4
1 Chemistry Department, Faculty of Science, University of Benghazi, Benghazi, Libya
2 Chemistry Department, Faculty of Science, Misurata University, Misurata, Libya
3 Department of Food Industries, Faculty of Agriculture, Misurata University, Misurata, Libya
4 Medical Laboratory Department, Faculty of Medical Technology, Misurata University, Misurata, Libya
Abstract
In the present study, silver complex with 8-hydroxyquinoline (8-HQ) was synthesized and characterized by FT IR and UV-VIS spectroscopic methods. The stoichiometric ratio, molar absorptivity, and formation constant were evaluated using UV-VIS spectrophotometric method. Both oxygen and nitrogen atoms from 8-HQ were included in the complex formation as stated by the spectral data. The UV-VIS spectrophotometric data indicated the formation of (1:1) complex with a formation constant of 4.2x108. Also, the antimicrobial effects of the ligand and the complex were examined in vitro. The diffusion style was used to investigate the antimicrobial activity on some chosen gram-positive and gram-negative bacteria. The ligand and its metal complex possessed antibacterial activity for particular bacteria such as Staphylococcus aureus and Proteus spp. The antimicrobial activity of the complex was higher with gram-negative bacteria. Meanwhile, the 8-HQ was found to have an appreciably higher capability to inhibit the growth of the gram-positive bacteria.
Graphical Abstract
Keywords
Introduction
8-hydroxyquinoline as a ligand is extensively used in coordination chemistry for the determination and extraction of metal ions [1]. It is also a significant compound which has the capability to coordinate with many metals as bidentate through nitrogen atom of quinolone ring and oxygen atom after deprotonation of hydroxyl group [2]. It has concerned extensive interest as a privileged structure and their derivatives have been examined for a wide extent of biological application [3].
In the latest years, specific regard has been assigned to the supramolecular coordination compounds established on 8-hydroxyquinoline and their derivatives [4]; specially, they are successfully applied to plan promising anti-cancer agents [5]. However, despite the continuous interest in novel 8-hydroxyquinoline as candidate anti-cancer agents, relatively few is recognized about their accurate mechanism of toxicity [6]. Chan et al. established that carbaldehyde derivative exhibited the highest in vitro cytotoxicity versus the human cancer cell lines [7].
Owing to the fast-rising phenomenon of drug resistance, investigation for modern and efficient antimicrobial agents is a worldwide importance [8-9]. Also, antibacterial agents which play a benefit controlling competing microorganism infection, are substantial. However, due to the emergence of antibiotic resistant bacterial strains, ordinary antibiotics are becoming less effective, resulting in incompetent treatment potency and considerable in health care costs [10-11].
Recently, appreciable researches have been assigned to study the antimicrobial and luminescent properties of silver (I) complexes [12-15]. The majority of these complexes have presented magnificent bactericidal properties [16]. They are also well recognized for their antimicrobial properties and excessively applied in the treatment of wounds and burn infections [17-20]. It has also displayed superb results towards various types of tumor cells, and this relies on the variety of ligand linked to the silver (I) ions [21-23].
The current paper aimed at the preparation and characterization of silver complex with 8-hydroxyquinoline ligand and investigation of its stoichiometry, molar absorptivity, and formation constant estimated by spectrophotometric method. Its antibacterial activity has also been investigated.
Experimental
Reagents and solvents
All Reagents and chemicals applied were of analytical grade and were utilized as received without any additional purification. 8-hydroxyquinoline (C9H7NO, ≥99.0) was purchased from Sigma-Aldrich. Silver nitrate (AgNO3, >99.8), acetonitrile (≥99.9%), and ethanol (≥99.9%) were from Supelco. Double distilled water was used for all the experiments.
Preparation of metal complex
The complex was synthesized by the method described in the literature [24]. A solvent of 99.9% ethanol and double distilled water were used to prepare the ligand and metal ion solutions, respectively. The concentration of ligand was 0.001 mol in 10 mL ethanol (0.145 g) and concentration of silver nitrate was also 0.001 mol in 10 mL double distilled water (0.170 g). The Prepared solutions (10 mL of each; the metal and ligand solutions) were mixed in a beaker and stirred by a magnetic stirrer for 30 minutes and then left in the dark for two days to precipitate a white complex. Then, they were filtered through Whatman filter paper No. 1 and stored in a desiccator.
Spectral analysis
The IR spectra of the ligand and synthesized silver complex were scanned from 650 to 4000 cm-1 using Perkin Elmer FT-IR Spectrometer. The UV-Visible spectra of them were recorded in the range from 200 to 400 nm using the Agilent Cary 60 UV-Vis Spectrophotometer. Spectra were recorded at 25 °C.
The stoichiometry of prepared complex (Ag-8-HQ) was determined by spectrophotometric method published previously by Joe and Yones [25]. A group of solutions with a constant concentration of 8-HQ of 2.00 ×10−6 mol/l were made, while Ag(I) concentrations were varied. According to the recorded results, the stoichiometric composition of the complex was determined and the values of Kf formation constant and molar absorptivity were calculated.
In Vitro antibacterial activity tests
The in vitro antimicrobial activities of the silver complex and 8-HQ were investigated using five strains of different bacteria (Staphylococcus aureus, Escherichia coli, Klebsiella spp., Proteus spp., and Salmonella spp.) The well diffusion technique was applied and it was according to the standard method of Thitilertdecha et al. [26]. Microorganism cell suspension at 0.5 McFarland standards was used. Two concentrations; 0.01 and 0.005 mg/mL, for both compounds were prepared by dissolving the silver complex and 8-HQ in acetonitrile and they were used for the assay. The sample volumes applied to the agar wells were approximately 20 μL of each solution. They were inoculated onto wells and were made in the spread plate culture of each microbial isolates; the plates were carried out in triplicates. All plates of the tested organisms were then allowed to incubate at 37 °C for overnight. After 18- 24 h of incubation, each solution was noted for inhibition zone for all isolates. The diameters of zone inhibitions were measured in millimeter scale (mm). As control, Ciprofloxacin antibiotic drug was used with a concentration of 5 mg/mL.
Source of microbial cultures
Cultures were obtained from department of Microbiology at Moafa Laboratory, Misuratua Libya. They included five strains of different bacteria, namely Staphylococcus aureus, Escherichia coli, Klebsiella spp., Proteus spp., and Salmonella spp. The bacterial cultures were maintained in their respective agar at around 5°C throughout the course of the study and used as stock cultures.
Results and Discussion
FT-IR spectra of HQ ligand and Ag-HQ complex
FT-IR spectra are the best prevailing means for detecting the functional groups in organic and inorganic compounds [27]. The FT-IR spectrum of the 8-hydroxyquinoline ligand (HQ) and its silver complex (Ag-HQ) are illustrated in Figure 1 and the frequency records are presented in Table 1.
In the ligand spectrum a band arose in the region of 3600 cm-1 that is attributed to the hydroxyl group present in 8-hydroxyquinoline aromatic ring. A peak rising at 3045 cm-1 is allocated to CH2 group vibration in the aromatic ring. A band rising in the region of 1600 cm-1 is allocated for C=N present in aromatic ring. A strong band rising in the range of 1300 cm-1 and 1400 cm-1 is allocated for C-N [24].
In the Ag complex spectrum of, a broad band seen was compared with its ligand in the region of 3370-3500 cm-1 owing to the coordination of the ligand with silver ions via the oxygen atom of phenolic group of 8-hydroxyquinoline and could also be because of the coordination of water molecules [28]. The band in the range of 1045 cm-1 is specified to the C-O-M in the corresponding metal complex. The band extends from 1330-1380 cm-1 due to C-N stretching vibrations of the metal complexes [24].
Figure 1. FT-IR spectra obtained for (a) Ag-HQ complex, b) HQ ligand
Table 1. FT-IR spectral records of the HQ ligand and its Ag complex
Electronic spectra of HQ ligand and Ag-HQ complex
The electronic spectra afford the details regarding the electronic structure of the ligand and its metal complex [29-31]. The UV-Visible spectrum obviously displays the electronic absorption of HQ ligand and its silver complex, which is revealed in Figure 2.
Figure 2. UV-Visible absorption spectra obtained for HQ and Ag-HQ complex
The electronic spectra of the ligand and the silver complex displayed various bands in the region 200-300 nm. The band releasing in the range of 200-250 nm is specified as π →π* transition of the ligand while the band at 250-300 nm is specified to chromophores metal complexes, which is a strong indication of the metal ion coordination with nitrogen and oxygen atoms existing in HQ ligand [32].
Metal complex stoichiometry and stability complex
The composition of Ag(I) complexes with HQ ligand was evaluated by the method of mole ratio [25]. A plot of absorbance versus mole ratio of the Ag(I) metal is constructed. As the complex formation is reasonably favorable; at a mole ratio consistent with the complex formation ratio, two straight lines of dissimilar slopes that intersect are obtained. Ideal mole-ratio plot is displayed in Figure 3. Formation constant of the complex can be estimated from the data in the curved portion of mole-ratio plot where the reaction is slightly complete. The complex molar absorptivity can also be estimated using the data obtained from the first straight line in Figure 3; therefore, in Figure 4, complex absorbance is plotted against concentration and consequently the complex molar absorptivity is calculated. The variables evaluated by mole ratio method; i.e. the complex stoichiometry, formation constant, and molar absorptivity values, are given in Table 2.
Figure 3. Mole ratio plot of the Ag-HQ complex
Figure 4. Calibration curve of the Ag-HQ complex
Table 2. The complex stoichiometry, formation constant, conductivity, and molar absorptivity values of the Ag-HQ complex
Antimicrobial investigations
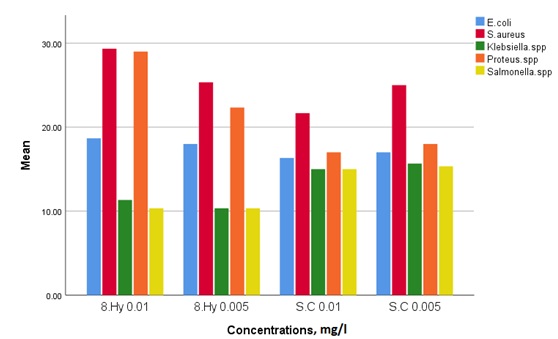
The obtained results disclosed several sensitivities of two compounds regarding the micro-organisms, as shown in Table 3. The results showed that all examined compounds have appropriate activities against all the tested micro-organisms. The silver complex and 8-HQ presented the highest activities against S. aureus as gram-positive bacteria and Proteus spp. as gram-negative bacteria. However, the 8-HQ displayed lower inhibition zones against Klebsiella spp. and Salmonella spp. The most efficient compounds between the two examined compounds against the five tested micro-organisms were silver complex (conc. of 0.005 mg/mL) and 8-HQ (conc. of 0.01 mg/mL) due to its broad spectrum of activity, as shown in Figure 5. Generally; by comparing the activities of the two compounds, a stronger activity of the silver complex compared with the 8-HQ with most Gram-negative bacteria was identified. Furthermore, statistical analysis on obtained results using ANOVA ONE WAY showed that there were significant differences.
Table 3. Analysis of variance (ANOVA) for antibacterial activity
Figure 5. Antibacterial activity of 8-HQ and synthesized complexes
Conclusion
Silver (I) complex with 8-HQ was prepared and characterized by FT-IR and UV-VIS spectroscopic techniques. The complexation between silver ion and the ligand was through the oxygen and nitrogen atoms of 8-HQ as indicated by IR spectra. Also, Ag(I) ions reacted with 8-HQ in a molar ratio of 1:1 (M:L), which was confirmed by mole ratio method. The formed complex was stable by the formation constant value (Kf=4.20x108), which were determined using spectrophotometric method. Finally, silver complex showed higher bacterial activity against gram-negative bacteria.
Acknowledgement
We are sincerely grateful to Misurata University and Moafa Laboratory for support of this work.
Disclosure statement
No potential conflict of interest was reported by the authors.
ORCID
Khaled M. Elsherif : 0000-0002-3884-1804